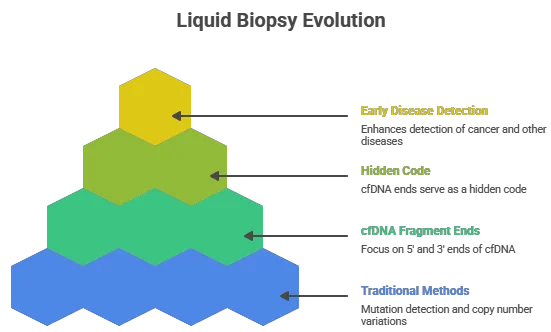
For decades, the field of liquid biopsy has focused primarily on three things: mutation detection, copy number variations, and methylation patterns. However, a revolutionary layer of biological information is now emerging from the shadows—one that does not change the genetic sequence but fundamentally alters how we read it.
In recent years, the field of liquid biopsy has undergone a transformative shift, moving beyond traditional methods of mutation detection, copy number variations, and methylation patterns. A new frontier has emerged, focusing on the intricate biology of cell-free DNA (cfDNA) fragment ends. This document explores the significance of the 5’ and 3’ ends of cfDNA fragments, revealing how they serve as a hidden code that can enhance early disease detection, including cancer, autoimmune diseases, and organ transplant rejection.
Introduction to cfDNA
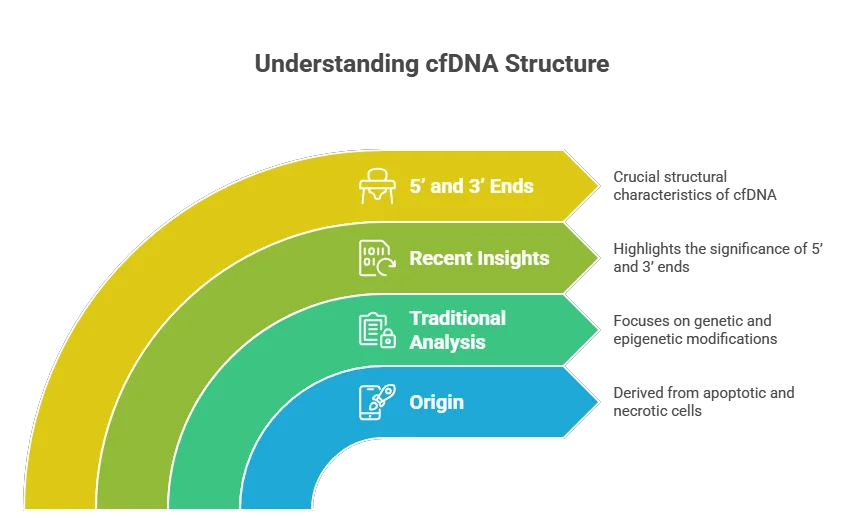
Cell-free DNA (cfDNA) refers to fragments of DNA that circulate freely in the bloodstream, originating from various sources, including apoptotic and necrotic cells. Traditionally, cfDNA analysis has centered on genetic alterations and epigenetic modifications. However, recent studies have illuminated the importance of the structural characteristics of cfDNA fragments, particularly their 5’ and 3’ ends.
The Importance of 5’ and 3’ Ends
The 5’ and 3’ ends of cfDNA fragments are not merely the starting and ending points of the DNA sequence; they carry critical information about the biological processes that produced them. These ends can reveal insights into the cellular environment, including the presence of specific enzymes and the conditions under which cells die.
Characteristics of cfDNA Fragment Ends
- Nucleotide Composition: The nucleotide sequences at the 5’ and 3’ ends can vary significantly based on the source of the cfDNA. For instance, fragments released from tumor cells may exhibit distinct patterns compared to those from healthy cells.
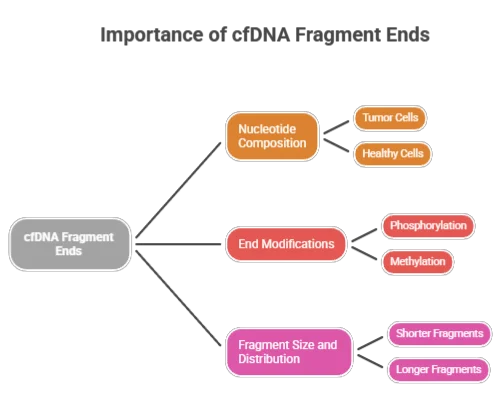
- End Modifications: Chemical modifications at the ends of cfDNA fragments, such as phosphorylation or methylation, can influence their stability and interaction with other biomolecules. These modifications can serve as biomarkers for various diseases.
- Fragment Size and Distribution: The size of cfDNA fragments can provide clues about the underlying pathology. For example, shorter fragments may be associated with certain types of cancer, while longer fragments might indicate other conditions.
Welcome to the world of cell-free DNA (cfDNA) fragment end biology.
Specifically, the precise characteristics of the 5’ and 3’ ends of cfDNA fragments are proving to be a hidden code—a cryptic lexicon written by dying cells. By learning to read this code, scientists are unlocking the potential for detecting cancer, autoimmune diseases, and organ transplant rejection months or even years earlier than conventional methods.
What Are “5’ and 3’ cfDNA Ends”?
To understand the code, we must briefly revisit molecular biology. Every DNA strand has directionality:
- The 5’ (five-prime) end typically carries a phosphate group.
- The 3’ (three-prime) end typically carries a hydroxyl group.
When a cell dies via apoptosis (programmed cell death) or necrosis, enzymes cleave the genomic DNA into fragments. These fragments—now cell-free DNA—float in the bloodstream. Crucially, the way the DNA is cut is not random.
Research published in Nature Genetics (2016) and Science Translational Medicine (2020) demonstrated that the 5’ and 3’ ends of cfDNA are not uniform. Instead, they exhibit specific end motifs (short sequences of 2–4 nucleotides at the termini) and end coordinates that reflect the activity of nucleases like DNASE1L3, DFFB, and DNASE1.
The Hidden Code: A New Paradigm
The concept of a “hidden code” in cfDNA fragment ends suggests that these structural features encode information that can be deciphered to understand disease states better. By analyzing the patterns and characteristics of the 5’ and 3’ ends, researchers can gain insights into:
- Tumor Dynamics: Changes in the cfDNA end characteristics can reflect tumor growth, metastasis, and response to treatment. This information can be invaluable for monitoring disease progression and therapeutic efficacy.
- Autoimmune Responses: In autoimmune diseases, the cfDNA profile may change due to the immune system’s attack on healthy tissues. Analyzing cfDNA ends can help identify these changes early, potentially leading to timely interventions.
- Organ Transplant Rejection: The presence of specific cfDNA end modifications can indicate whether an organ transplant is being rejected. Early detection of rejection can significantly improve patient outcomes.
The Hidden Code: Why Ends Matter More Than Mutations
While mutations tell you what is wrong, the ends of cfDNA tell you who is dying, how they are dying, and where in the body it is happening.
1. Tissue-of-Origin Mapping
Different cell types express different nucleases. Liver cells, for example, utilize specific enzymes that leave a unique “signature” on the 5’ and 3’ ends of cfDNA. By sequencing the ends of thousands of fragments, researchers can deconvolute the mixture of DNA in the blood. A disruption in normal end patterns—such as an overrepresentation of fragments ending with “C-C-A” motifs—may point to hepatocyte death from a tumor or hepatitis.
2. Fragmentomics: The Shape of the End
The field of fragmentomics analyzes the size and end properties of cfDNA. Healthy individuals primarily have fragments that are ~167 base pairs long—wrapped around a nucleosome. But cancerous cells alter the enzymatic landscape. Tumors often produce cfDNA with different end coordinates, resulting in shorter, jagged fragments. The specific base at the 5’ end (e.g., a predominance of cytosine vs. thymine) serves as a fingerprint for the disease state.
3. Early Detection of Fragmentation Disorders
In autoimmune diseases like systemic lupus erythematosus (SLE), the body fails to clear debris from dead cells. This results in an accumulation of cfDNA with blunted or modified ends that are normally degraded quickly. By quantifying the ratio of “smooth” ends (apoptotic) to “ragged” ends (necrotic), clinicians can diagnose disease flares weeks before clinical symptoms appear.
Methodologies for Analyzing cfDNA Ends
To harness the potential of cfDNA fragment end analysis, several methodologies have been developed:
- Next-Generation Sequencing (NGS): NGS allows for comprehensive analysis of cfDNA, enabling researchers to examine the nucleotide composition and modifications at the 5’ and 3’ ends in great detail.
- Bioinformatics Tools: Advanced bioinformatics algorithms can analyze large datasets generated by NGS, identifying patterns and correlations that may indicate disease presence or progression.
- Liquid Biopsy Platforms: Emerging liquid biopsy technologies are being designed to specifically target cfDNA ends, facilitating rapid and accurate disease detection.
Clinical Implications
The ability to decode the 5’ and 3’ ends of cfDNA fragments holds significant promise for clinical applications:
- Early Detection: By identifying disease-specific patterns in cfDNA ends, clinicians may be able to detect conditions like cancer or autoimmune diseases at much earlier stages than currently possible.
- Personalized Medicine: Understanding the unique cfDNA profiles of individual patients can lead to more tailored treatment strategies, improving outcomes and minimizing side effects.
- Monitoring Disease Progression: Regular analysis of cfDNA can provide ongoing insights into disease status, allowing for timely adjustments in treatment plans.
How Machine Learning Decodes the Termini
The human genome contains approximately 3 billion base pairs. The possible combinations of 5’ and 3’ ends are astronomical. This is where artificial intelligence (AI) and deep learning have become indispensable.
Modern liquid biopsy assays now employ end-motif profiling. An AI model is trained on thousands of cfDNA samples from healthy donors, cancer patients, and transplant recipients. The model learns to recognize subtle shifts in the frequency of specific 4-base end motifs (e.g., “CATA” at the 5’ end and “TGTA” at the 3’ end).
A landmark study from the University of Chicago (2022) demonstrated that 5’ and 3’ end signatures alone could detect early-stage hepatocellular carcinoma (HCC) with 91% sensitivity—without ever looking for a known mutation. The code was hiding in the ends.
Clinical Applications on the Horizon
| Condition | Hidden Code Signature | Clinical Utility |
|---|---|---|
| Early Cancer | Shortened fragments (<100 bp) with non-canonical 5’ ends (e.g., high “A” or “T” starts) | Screening for pancreatic and ovarian cancers months before CT scans. |
| Organ Transplant Rejection | Donor-derived cfDNA with distinct 3’ end preferences (e.g., “G” ending) | Non-invasive monitoring to replace risky biopsies. |
| Myocardial Infarction | Sharp increase in 5’ hydroxylated ends (necrotic signatures) from cardiomyocytes | Emergency diagnosis vs. benign chest pain. |
| Pregnancy Complications | Aberrant 5’ methylation patterns on placental cfDNA ends | Early prediction of preeclampsia. |
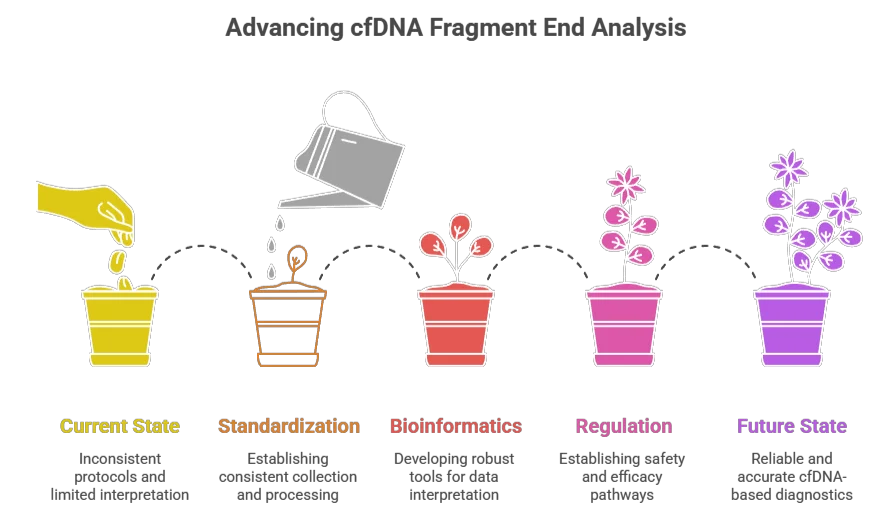
Challenges and the Road Ahead
Despite its promise, decoding the 5’ and 3’ cfDNA ends is not yet routine. Current challenges include:
- Library Preparation Bias: Many sequencing methods introduce artificial ends during DNA extraction. Single-strand library prep is required to preserve true biological ends.
- Bioinformatics Complexity: Storing and analyzing end-coordinate data requires massive computational resources and novel algorithms.
- Normal Variation: Diet, exercise, and circadian rhythms subtly alter end-motif frequencies. Defining a “normal” baseline is an ongoing effort.
Nevertheless, companies like Delfi Diagnostics, Grail, and Freenome are quietly integrating end-motif analysis into their next-generation platforms. The era of “end-omics” has begun.
Conclusion: Reading the Final Breath of Cells
Every day, billions of cells in your body die and release their DNA into your blood. The 5’ and 3’ ends of those fragments are not random noise—they are a history book. They record the enzyme that killed the cell, the tissue it came from, and whether the death was programmed or pathological.
By learning to read this hidden code, we are moving beyond the “what” of disease genetics to the “when” and “where.” In the coming decade, a simple blood test analyzing the ends of cfDNA may become the standard for annual check-ups, catching cancer, autoimmune flares, and organ damage before symptoms ever appear.
The code has been hiding in plain sight—at the ends.
Disclaimer: This article is for informational purposes and does not constitute medical advice. Always consult a healthcare provider for diagnostic decisions.