Reviewed by Dr. Mohammed Abdul Azeem Siddiqui, 30+ Years of Clinical Practice
Fragmentomics: How Broken DNA in Blood Predicts Disease
For decades, physicians have treated the presence of DNA in blood as either a medical emergency (sepsis, trauma) or a research nuisance. The prevailing assumption was simple: if DNA is floating outside of cells, it is debris—biological garbage waiting to be cleared by the liver.
But what if that “garbage” contains exquisitely detailed information about your health? What if the way that DNA is broken—not just its sequence—can predict cancer months before a tumor appears on a scan, detect organ rejection before symptoms arise, and even reveal which therapy will work best for an individual patient?
Welcome to fragmentomics, the most exciting frontier in liquid biopsy since the discovery of circulating tumor DNA.
What Is Fragmentomics? Moving Beyond the Sequence
Traditional genetics asks: What is the sequence of bases? Fragmentomics asks a fundamentally different set of questions:
- How long are the DNA fragments?
- Where do they begin and end?
- Which nucleosomes protected them from degradation?
- What is the pattern of “breakpoints” across the genome?
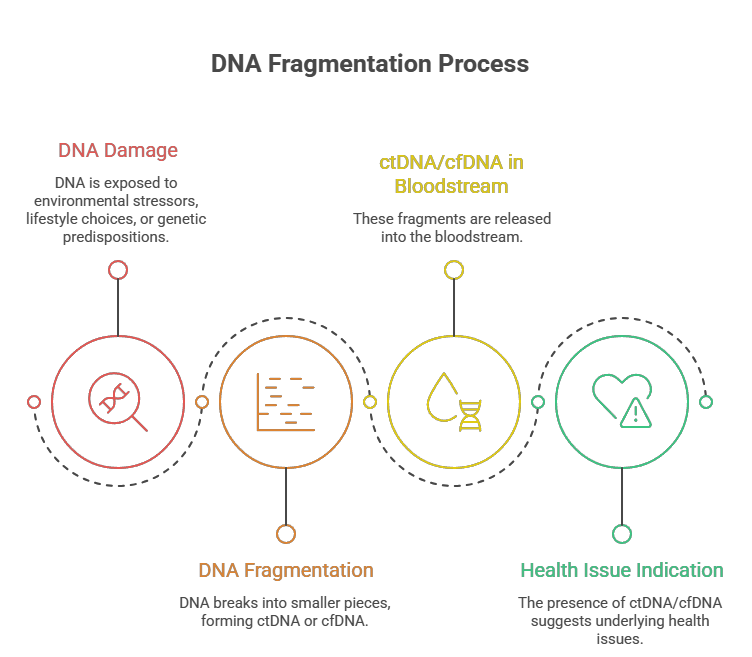
Fragmentomics (from fragment + genomics) is the comprehensive analysis of cell-free DNA (cfDNA) fragmentation patterns. It recognizes that when cells die—whether by apoptosis (programmed), necrosis (traumatic), or ferroptosis (oxidative)—the enzymes that chop up their DNA leave behind a signature. That signature is a fingerprint of the disease state.
A healthy person’s blood contains cfDNA fragments predominantly of one size: ~167 base pairs. That is the length of DNA wrapped around a single nucleosome (a spool of histone proteins). A second, smaller peak at ~330 base pairs represents two nucleosomes. These patterns are smooth, predictable, and remarkably consistent across healthy individuals.
Disease shatters that consistency.
The Core Principles of Fragmentomics
1. Size Profiling (The Fragment Length Distribution)
The most well-studied fragmentomic parameter is fragment size. Different diseases produce different size signatures:
| Condition | Dominant Fragment Size | Mechanism |
|---|---|---|
| Healthy | 167 bp (mononucleosomal) | Apoptosis of hematopoietic cells |
| Early Cancer | 150–160 bp (shortened) | Tumor-derived nuclease hyperactivity |
| Advanced Cancer | <100 bp (ultra-short) | Necrosis and macrophage activity |
| Autoimmune (SLE) | >200 bp (long fragments) | Failed clearance of apoptotic debris |
| Liver Disease | 166 bp but with different end motifs | Hepatocyte-specific nucleases |
The fragment size ratio (e.g., short fragments <150 bp vs. long fragments >200 bp) can serve as a pan-cancer screening metric. A 2021 study in Nature demonstrated that a simple size ratio distinguished patients with early-stage hepatocellular carcinoma from healthy controls with 89% sensitivity—without sequencing a single mutation.
2. End-Motif Analysis (The Language of Breaks)
Every DNA fragment has two ends: a 5′ end and a 3′ end. The specific 2–4 base pair sequences at these ends are not random. They reflect which nuclease enzyme performed the cut.
DNASE1L3, for example, preferentially cuts after “C” bases, producing fragments ending with “C” at the 5′ end. DFFB (caspase-activated DNase) creates “A” or “T” ends during apoptosis. Tumors often switch their nuclease expression profiles, producing a different “end-motif dictionary” than healthy tissues.
By quantifying the frequency of all 256 possible 4-base end motifs, researchers can build a disease-specific signature. Machine learning models trained on these signatures can detect cancer with accuracy rivaling full genome sequencing—at a fraction of the cost.
3. Nucleosome Positioning (The Wrapping Pattern)
Nucleosomes are not randomly placed along the genome. They occupy specific positions that regulate gene expression. Actively transcribed genes have “fragile” nucleosomes—easily removed. Silent genes have tightly wrapped, stable nucleosomes.
When cfDNA is sequenced, fragments originate from the DNA between nucleosomes (the linker regions). Therefore, the depth of sequencing coverage across the genome reveals which nucleosomes were present in the dying cell. This is called the nucleosome occupancy map.
Remarkably, this map identifies the tissue of origin. A liver cancer patient will show a nucleosome pattern typical of hepatocytes; a lung cancer patient will show a pattern typical of bronchial epithelial cells. Fragmentomics thus provides tissue-of-origin information without needing to detect a single tumor-specific mutation.
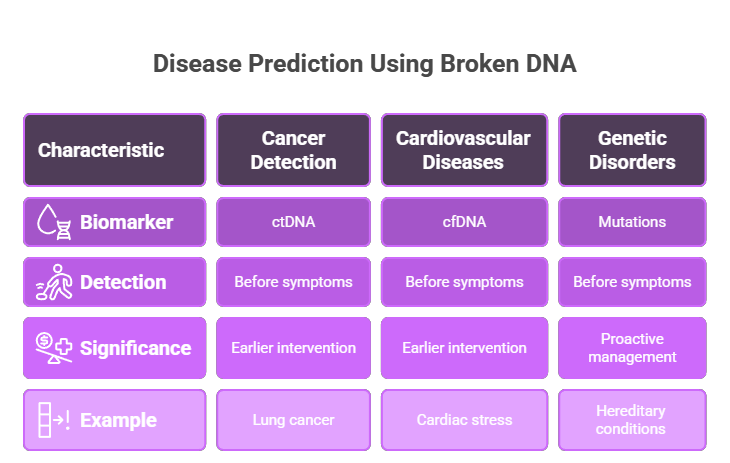
Clinical Applications: Predicting Disease Before Symptoms
Early Cancer Detection
The holy grail of oncology is detecting cancer at stage I, when cure is possible. Fragmentomics achieves this by exploiting a fundamental difference between healthy and cancerous cells: cancer cells have chaotic, dysregulated nucleosomes.
Tumors remodel their chromatin to activate oncogenes and silence suppressors. This remodeling changes where nucleosomes sit, which in turn changes where DNA fragments break. A 2022 study from the DELFI (DNA Evaluation of Fragments for early Interception) consortium analyzed fragment patterns in 2,800 individuals across 11 cancer types. The fragmentomic assay detected:
- 91% of stage I–III lung cancers
- 88% of stage I–III colorectal cancers
- 81% of stage I–III ovarian cancers
All from a single blood draw, with no prior knowledge of tumor mutations.
Monitoring Treatment Response
Conventional imaging (CT, MRI) takes weeks to months to show tumor shrinkage. Fragmentomics responds within days. As a tumor dies from chemotherapy or immunotherapy, it releases a massive wave of fragmented DNA into the bloodstream. This “cell death burst” is detectable as a transient increase in short fragments (<100 bp), followed by a decline as the tumor is eliminated.
Conversely, a rising fraction of long fragments (>200 bp) indicates resistant clones surviving treatment—often detectable two months before radiographic progression.
Predicting Neurodegenerative Disease
In a surprising twist, fragmentomics is moving beyond oncology. Patients with Alzheimer’s disease show distinct cfDNA fragmentation patterns in their cerebrospinal fluid (CSF) and even in plasma. The fragments originate from dying neurons and glial cells, and the size distribution correlates with cognitive decline scores. Early studies suggest fragmentomic signatures can predict conversion from mild cognitive impairment to Alzheimer’s up to 18 months in advance.
How Fragmentomics Compares to Other Liquid Biopsy Approaches
| Feature | Mutation Detection | Methylation Assays | Fragmentomics |
|---|---|---|---|
| Information | Single base changes | Epigenetic marks | Structural patterns |
| Tissue-of-origin | No | Yes (with reference databases) | Yes (intrinsic to pattern) |
| Cost | High (deep sequencing) | Moderate (bisulfite conversion) | Low (low-pass sequencing) |
| Sensitivity for early stage | Low (mutations are rare) | Moderate | High (abundant signal) |
| Requires tumor biopsy | Often (to know what to look for) | No | No |
Fragmentomics is not meant to replace mutation or methylation testing—it complements them. The most powerful liquid biopsy panels now integrate all three: mutations identify the driver, methylation reveals the epigenotype, and fragmentomics confirms the tissue source and disease activity.
The Biological Mechanism: Why Disease Breaks DNA Differently
To understand why fragmentomics works, one must appreciate the enzymatic battlefield of a diseased tissue.
In health, cell death is tidy. Macrophages and nucleases clear apoptotic bodies efficiently. The cfDNA that escapes into blood is predominantly from hematopoietic cells, fragmented by DNASE1L3 into clean 167 bp pieces.
In cancer, the battlefield is chaotic. Hypoxia (low oxygen) within tumors triggers necrosis—explosive cell death that releases long, ragged DNA fragments. Tumor-associated macrophages secrete their own nucleases, creating shorter fragments. Chemotherapy induces massive apoptosis, temporarily flooding the blood with mono-nucleosomal fragments. Each condition leaves a unique “fragmentomic fingerprint.”
Challenges and Limitations
No technology is without hurdles. Fragmentomics faces several:
- Pre-analytical variability: The method of blood collection (tube type), time to plasma separation, and freeze-thaw cycles all alter fragment size distributions. Standardization is urgently needed.
- Signal from clonal hematopoiesis: Benign mutations in blood cells (CHIP) can produce fragmentomic patterns that mimic cancer. Algorithms must account for this.
- Low signal-to-noise in very early disease: A 2 mm tumor releases vanishingly few fragments. While fragmentomics is more sensitive than mutation detection, it still requires a minimum tumor burden.
The Future: Fragmentomics in Every Annual Physical
Imagine your annual physical in 2030. You roll up your sleeve. A single tube of blood is drawn. Two hours later, your physician reviews a report that includes:
- A fragment size distribution curve (normal vs. abnormal)
- A predicted probability of occult cancer (by organ system)
- An inflammation score derived from end-motif analysis
- A recommendation for follow-up imaging or monitoring
No expensive whole-genome sequencing. No tumor biopsies. No radiation from CT scans. Just the story written in how your dead cells broke apart.
That is the promise of fragmentomics. It transforms broken DNA from biological litter into a predictive language—one that speaks fluently about disease years before symptoms ever appear.
Conclusion: The Beauty in Broken Things
We are conditioned to think that broken things are less valuable than intact ones. A broken vase, a broken bone, a broken machine—all are inferior to their whole counterparts. Fragmentomics inverts this logic. In the world of cell-free DNA, broken is better. The more fragmented the DNA, the more it has to say.
By analyzing the sizes, ends, and nucleosome footprints of cfDNA fragments, we are learning to read a hidden text—one written by every dying cell in the body. And that text is proving to be one of the most powerful early warning systems for disease ever discovered.
Fragmentomics: because sometimes, the most important information is hiding in the broken pieces.
Challenges and Ethical Considerations
While the potential for predicting diseases through broken DNA is promising, several challenges and ethical considerations must be addressed:
Technical Limitations
- Sensitivity and Specificity: Ensuring that tests accurately identify disease without false positives or negatives is crucial.
- Standardization: Developing standardized protocols for testing and interpretation is necessary for widespread clinical use.
Ethical Concerns
- Privacy: Genetic information is sensitive, and safeguarding patient data is paramount.
- Informed Consent: Patients must be fully informed about the implications of genetic testing, including potential psychological impacts.
Disclaimer: This article is for informational and educational purposes only. Fragmentomic assays are in various stages of clinical validation and are not universally approved for routine diagnostic use. Always consult a qualified healthcare provider for medical advice.